
Bona fide peaks will have multiple overlapping reads with offsets, while samples with only PCR duplicates will stack up perfectly without offsets. Try to discriminate via genome browser of your non-deduplicated data.
Although it was developed for the detection of transcription factor binding sites it is also suited for larger regions. The MACS algorithm captures the influence of genome complexity to evaluate the significance of enriched ChIP regions. MACS2Ī commonly used tool for identifying transcription factor binding sites is named Model-based Analysis of ChIP-seq (MACS). An example of this is the binding properties of PolII, which binds at promotor and across the length of the gene resulting in mixed signals (narrow and broad). There are also ‘mixed’ binding profiles which can be hard for algorithms to discern. Narrow peaks are easier to detect as we are looking for regions that have higher amplitude and are easier to distinguish from the background, compared to broad or dispersed marks. histone modifications that cover entire gene bodies) or narrow peaks (i.e. ChIP-seq analysis algorithms are specialized in identifying one of two types of enrichment (or have specific methods for each): broad peaks or broad domains (i.e. NOTE: Our dataset is investigating two transcription factors and so our focus is on identifying short degenerate sequences that present as punctate binding sites. Note that in this Session the term ‘tag’ and sequence ‘read’ are used interchangeably. One of the more commonly used peak callers is MACS2, and we will demonstrate it in this session. There are various tools that are available for peak calling. Image source: Wilbanks and Faccioti, PLoS One 2010 The distributions of these groups are then assessed using statistical measures and compared against background (input or mock IP samples) to determine if the site of enrichment is likely to be a real binding site. The 5’ ends of the selected fragments will form groups on the positive- and negative-strand. Peak calling, the next step in our workflow, is a computational method used to identify areas in the genome that have been enriched with aligned reads as a consequence of performing a ChIP-sequencing experiment.įor ChIP-seq experiments, what we observe from the alignment files is a strand asymmetry with read densities on the +/- strand, centered around the binding site. List and describe the output files from MACS2.Describe the parameters involved in running MACS2.
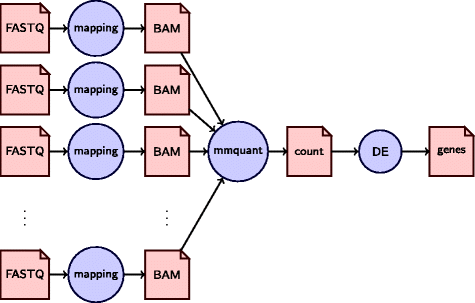
Describe the different components of the MACS2 peak calling algorithm.Peak calling with MACS2 Intro to ChIPseq using HPC View on GitHubĬontributors: Meeta Mistry, Radhika KhetaniĪpproximate time: 80 minutes Learning Objectives Peak calling with MACS2 | Introduction to ChIP-Seq using high-performance computing Skip to the content.

